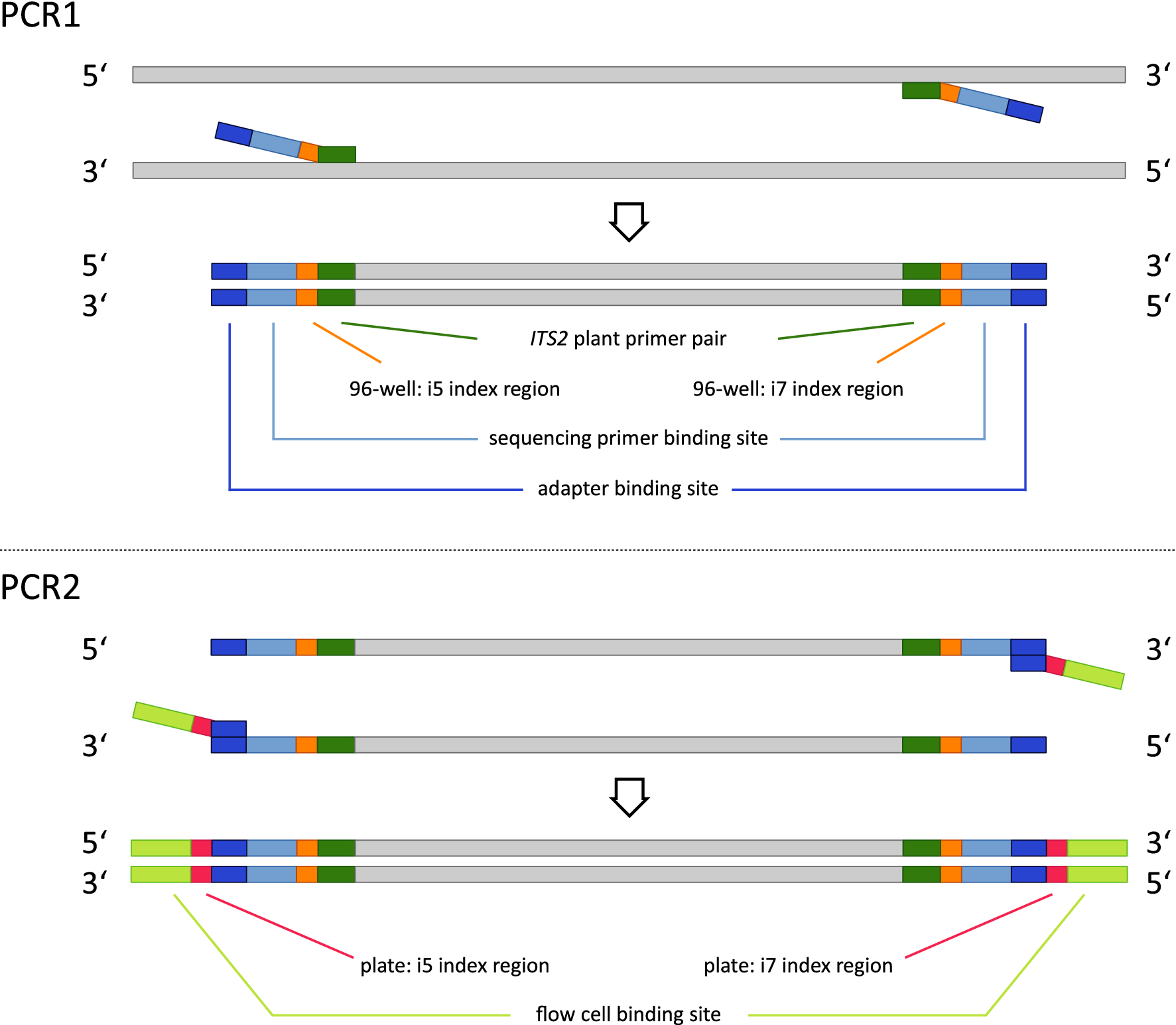
Handling of targeted amplicon sequencing data focusing on index hopping and demultiplexing using a nested metabarcoding approach in ecology | Scientific Reports

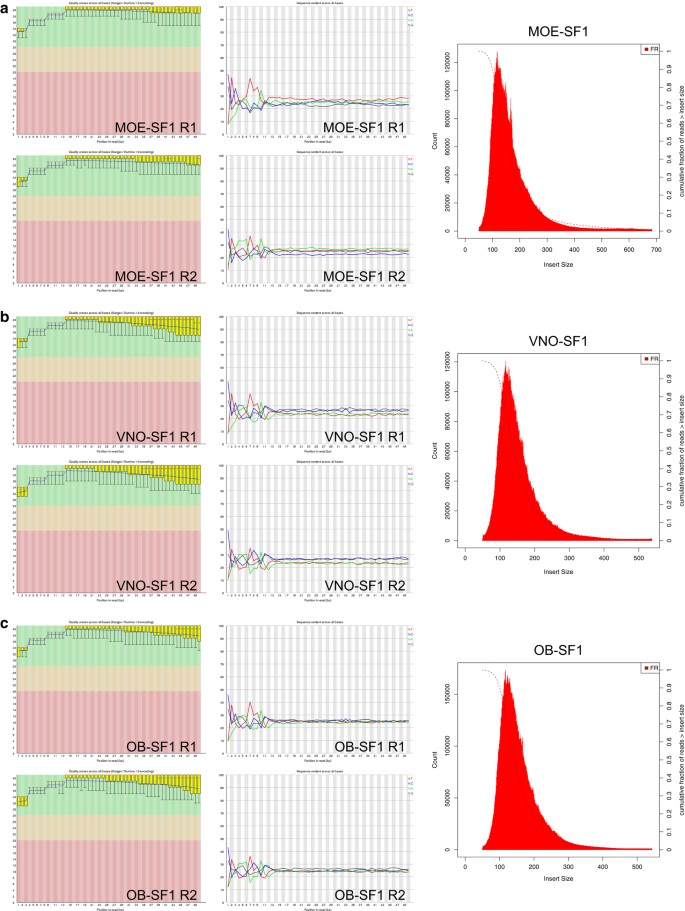
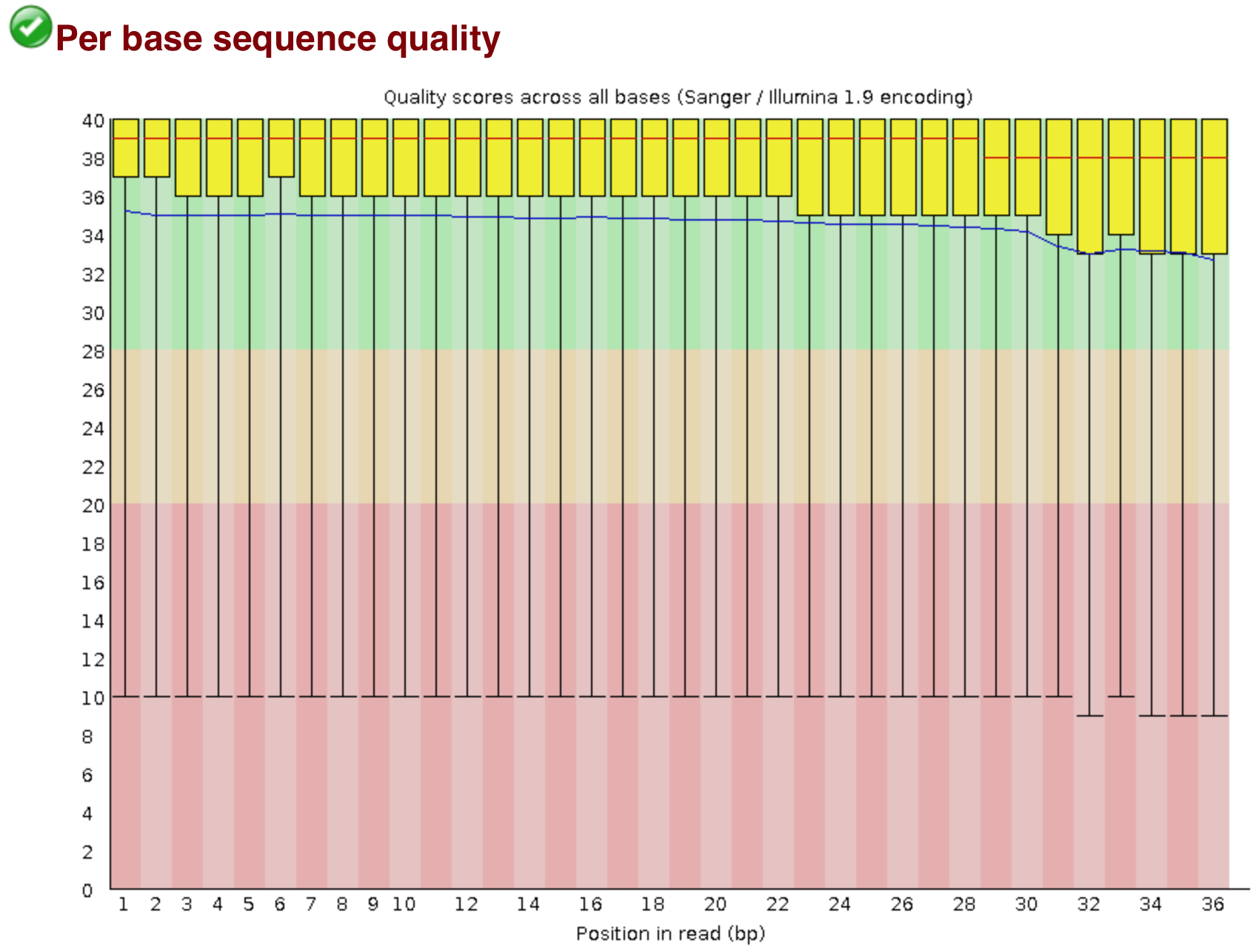
ClinQC quality control report generated by FASTQC. a Per base sequence... | Download Scientific Diagram

Biomolecules | Free Full-Text | High-Throughput Identification of Adapters in Single-Read Sequencing Data
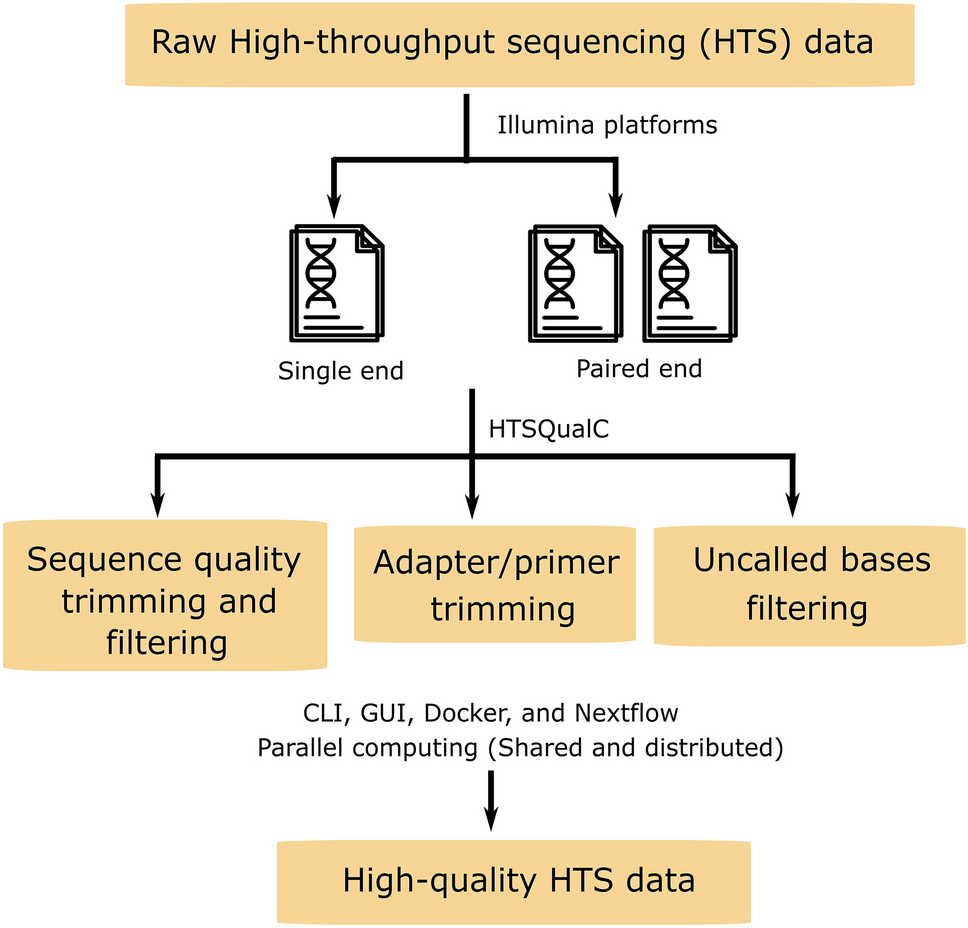
HTSQualC is a flexible and one-step quality control software for high-throughput sequencing data analysis | Scientific Reports

Statistical guidelines for quality control of next-generation sequencing techniques | Life Science Alliance
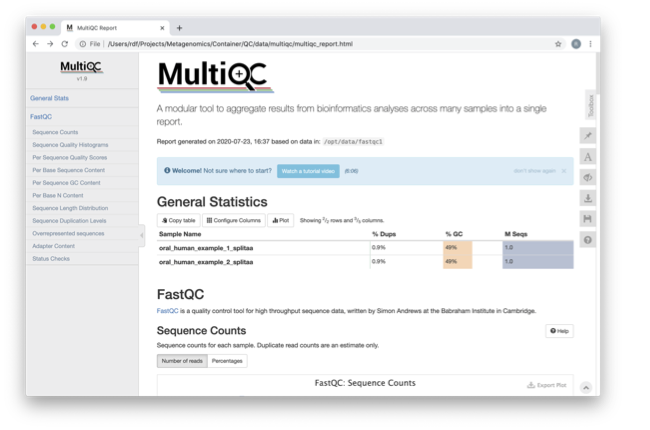
![PDF] FQC Dashboard: integrates FastQC results into a web-based, interactive, and extensible FASTQ quality control tool | Semantic Scholar PDF] FQC Dashboard: integrates FastQC results into a web-based, interactive, and extensible FASTQ quality control tool | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/eacf7e5551a8cea85278c2a746714f20258664ea/2-Figure1-1.png)
PDF] FQC Dashboard: integrates FastQC results into a web-based, interactive, and extensible FASTQ quality control tool | Semantic Scholar
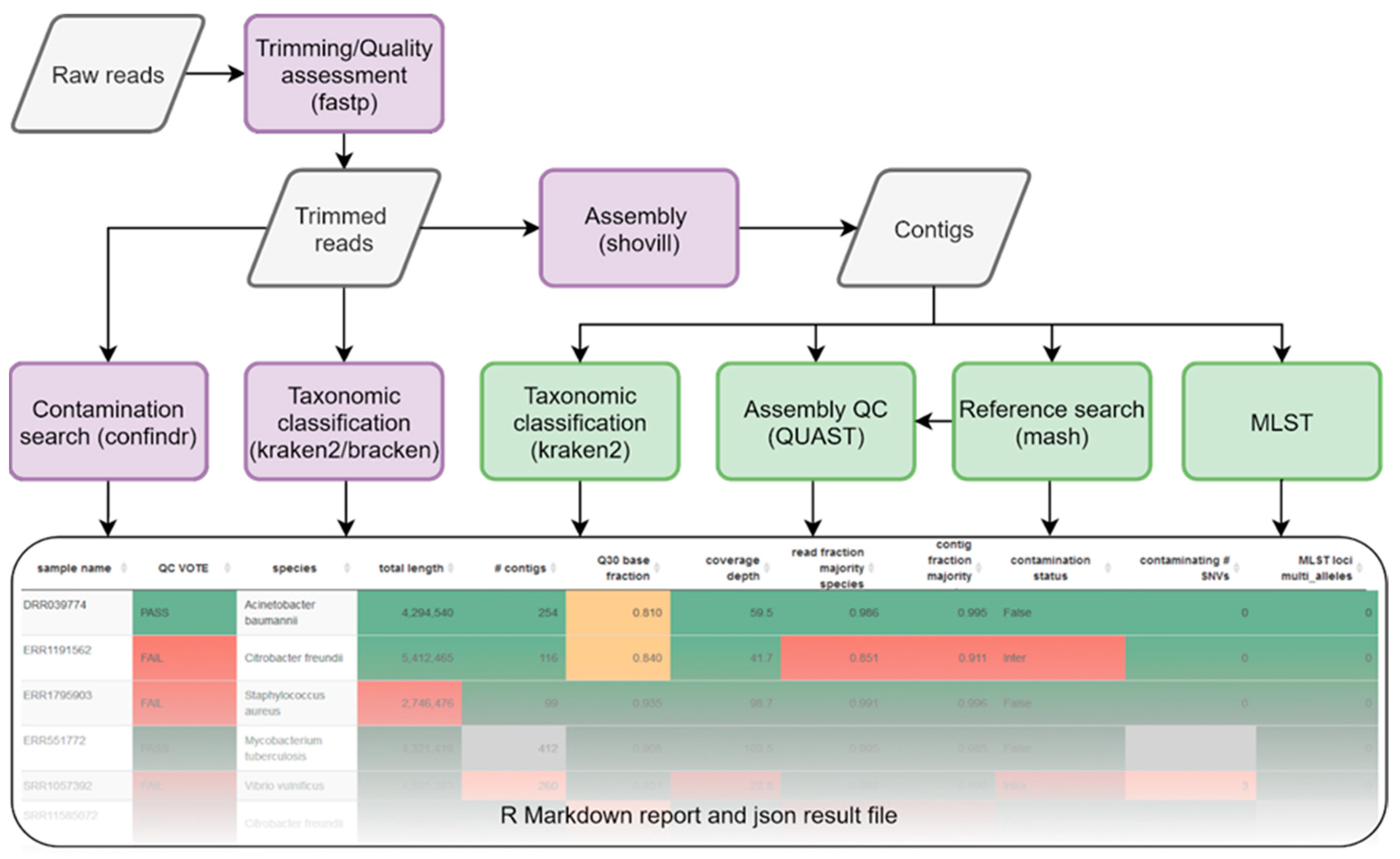
Genes | Free Full-Text | Species-Specific Quality Control, Assembly and Contamination Detection in Microbial Isolate Sequences with AQUAMIS
GitHub - natir/fastQt: FastQC port to Qt5: A quality control tool for high throughput sequence data.